 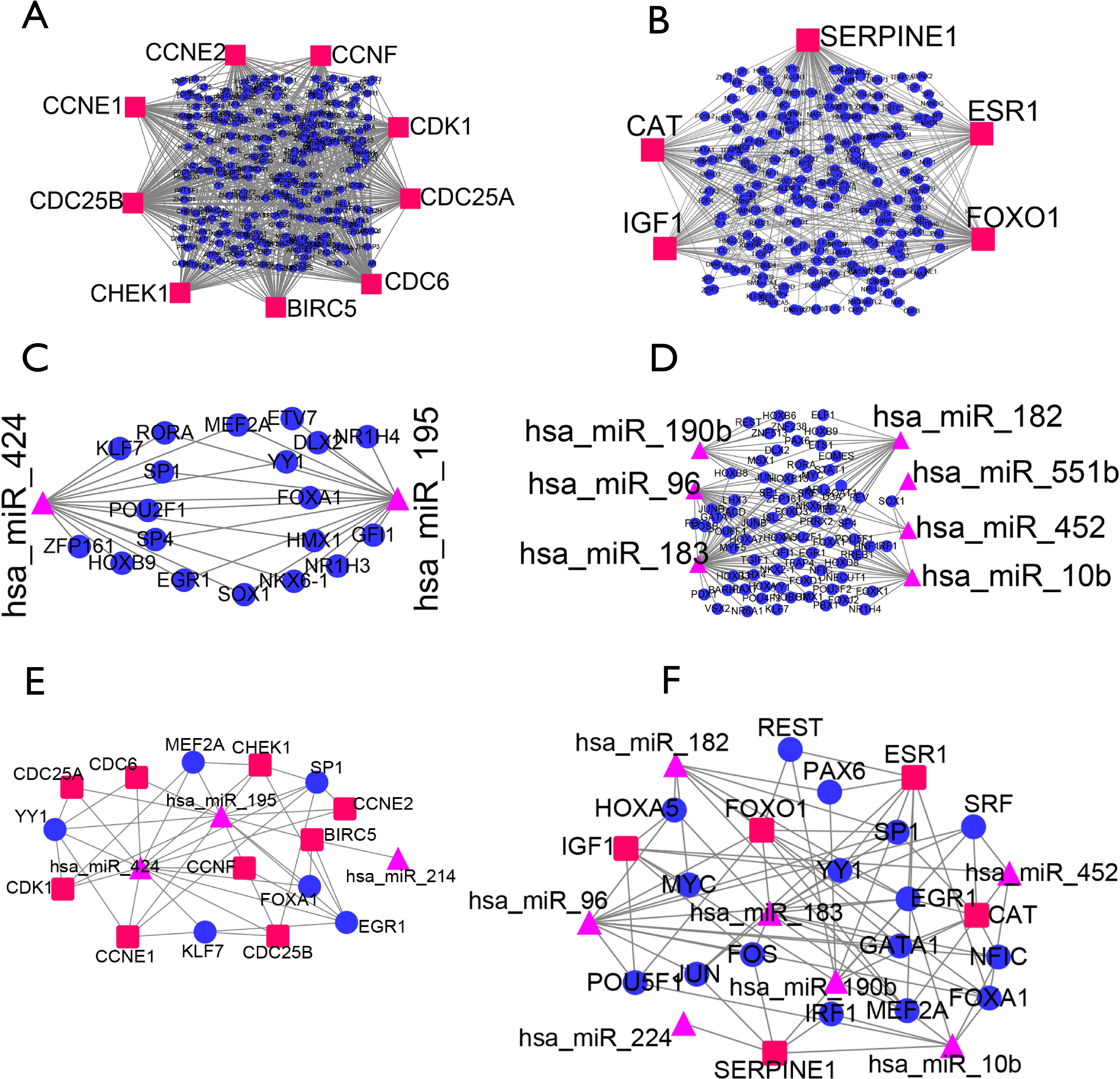
Profiling of TFBS motifs across miRNA promoter regions showed that binding site motifs for TCP, AP2/ERF, GATA, NF-YB, DOF, B3, bZIP, trihelix, ZF-HD, bHLH and Dehydrin transcription factor families are abundant in the Musa acuminata genome. Gene ontology of the 20 transcription factors among the predicted miRNA targets, showed predominance for auxin-activated signalling and developmental processes. Following this, banana root degradome data was used to confirm miRNA targets and the transcription factor targets were placed into a predicted network together with their targeting miRNA using cytoscape. A total of 183 mature miRNAs, comprising 144 orthologous miRNA and 39 Musa-specific miRNA were predicted. In this study, genome-wide annotation of miRNA in the most recently published banana genome sequence was used to predict miRNA promoter regions and to map TFBS of miRNA genes. Plant miRNA have been reported both to target and to be regulated by transcription factors, however, genomic distribution of miRNA, transcription factor targets for miRNA, transcription factor binding sites (TFBS) of miRNA promoters and their regulatory networks have not been systematically mapped in banana. 42(Database issue): D654-9).MicroRNA (miRNA) are important regulators of gene expression. #CYTOSCAPE TRANSCRIPTION FACTOR PLUS#Plus a huge database of transcriptional units. It uses a library of positional weight matrices from TRANSFAC® Public 6.0 along with the site alignments associated with these matrices.ĭOOR2 - Database of pr Okaryotic Ope Rons - offers high-performance web service for online operon prediction on user-provided genomic sequences and, an intuitive genome browser to support visualization of user-selected data. 
It uses a library of positional weight matrices from TRANSFAC® Public 6.0 and, P-Match - a program for predicting transcription factor binding sites (TFBS) in DNA sequences that combines pattern matching and weight matrix approaches. Provides access to programs including Match which is a weight matrix-based program for predicting transcription factor binding sites (TFBS) in DNA sequences. #CYTOSCAPE TRANSCRIPTION FACTOR PROFESSIONAL#TRANSFAC -offers academic and non-profit organizations free access to TRANSFAC® non-professional 2005 version with much reduced functionality and content compared to our professional database. TFmiR - for deep and integrative analysis of combinatorial regulatory interactions between transcription factors, microRNAs and target genes that are involved in disease pathogenesis. The predictions can be performed by four different methods (CONREAL-, LAGAN-, MAVID- and BLASTZ-based) and results can be compared to each other. Excluding up to 95% false positive transcription factor binding sites predictions while maintaining high sensitivity of the search.ĬONREAL - allows identification of transcription factor binding sites (TFBS) that are conserved between two sequences. RVista (Comparative Genomics Center, Lawrence Livermore National Laboratory, U.S.A.) - High-throughput discovery of functional regulatory elements in sequence alignments. 38: D822-D872) or here for related site.ĭBD: Transcription factor prediction database (Gesellschaft für Biotechnologische Forschung mbH (GBF), Braunschweig, Germany) ( Reference: D. PlnTFDB - Plant Transcriptional Factor Database - allows BLAST searching ( Reference: P. PLACE (National Institute of Agrobiological Sciences, Japan) - Plant cis-acting regulatory DNA elements. Tfsitescan (Institute for Transcriptional Informatics, Pittsburgh, U.S.A.) - This tool is intended for promoter sequence analysis and works best with sequences of ~500 nt. Transcription factor binding sites: several sites are available but I have not been particular impressed with the results when analyzing prokaryotic sequences 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
